Execution Flow by Request
This page provides an overview of the rule execution flow tailored to different request specified in the job configuration file of NovaScope.
Each request option triggers a specific set of rules. Thus, below provides a rulegraph for each request option to outline the triggered rules and their interdependencies, detailing distinct processing paths.
Info
The visual guides below are constructed from a baseline scenario where only the initial input 1st-seq and 2nd-seq FASTQ files are present, with no prior processing or intermediate files generated.
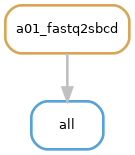
Request "sbcd-per-flowcell"¶
-
Description: The
"sbcd-per-flowcell"option initiates the generation of a spatial barcode map specific to a flow cell. Consequently, it triggers the execution of Rulefastq2sbcd. -
Rule Graph:
Request "sbcd-per-chip"¶
-
Description: The
"sbcd-per-chip"option requests a spatial barcode map specific to a given section chip and a plot of the distribution of spatial barcodes, triggering the execution of Rulesbcd2chipand its prerequisite rule:fastq2sbcd. -
Rule Graph:

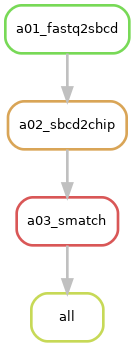
Request "smatch-per-chip"¶
-
Description: For each pair of input 2nd-seq FASTQ files associated with a given chip, the
"smatch-per-chip"option demands two files, including a file of spatial barcodes matched to the 2nd-seq reads alongside its summary metrics, and an image depicting the spatial distribution of those matched barcodes. This request prompts the execution of Rulesmatchand its prerequisite rules:sbcd2chipandfastq2sbcd. -
Rule Graph:
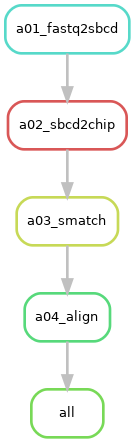
Request "align-per-run"¶
-
Description: The
"align-per-run"option request a Binary Alignment Map (BAM) from the alignment as well as the digital gene expression matrices (DGEs) for Gene, GeneFull, splice junctions (SJ), and Velocyto. It provokes the execution of Rulealignand its prerequisite rules includingsmatch,sbcd2chip, andfastq2sbcd. -
Rule Graph:
Request "sge-per-run"¶
-
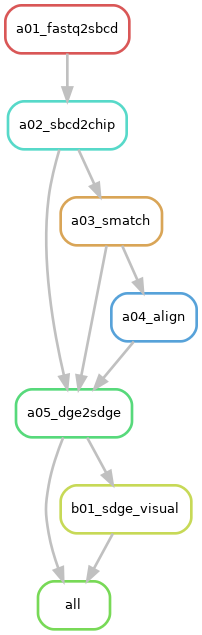
Description: The
"sge-per-run"option inquires: 1) a spatial digital gene expression matrix (SGE) encompassing all genomic features with two plots visualizing the distribution of the aligned spatial barcodes; 2) visualization for genes of interest provided in the job configuration file. It requests the execution of Ruledge2sdgeandgene_visual, alongside the prerequisite rules:align,smatch,sbcd2chip, andfastq2sbcd. -
Rule Graph:
Request "transcript-per-unit"¶
-
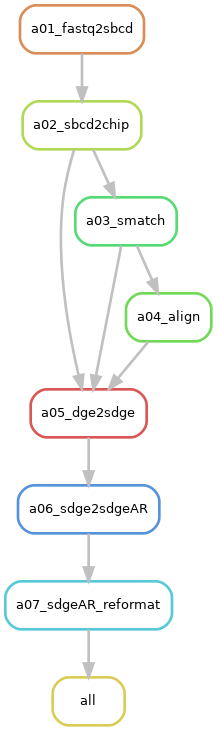
Description: The
"transcript-per-unit"option requests a SGE in a FICTURE-compatible format. It requests the execution of RulesdgeAR_reformat, alongside the prerequisite rules:sdge2sdgeAR,dge2sdge,align,smatch,sbcd2chip, andfastq2sbcd. -
Rule Graph:
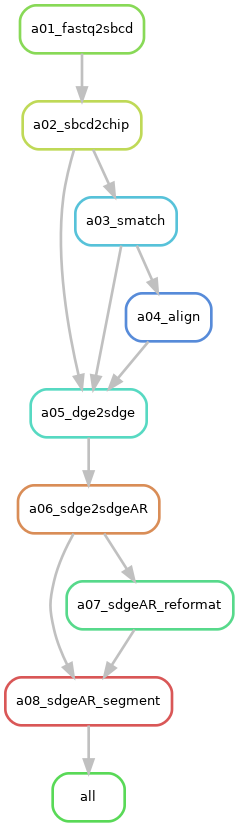
Request "segment-per-unit"¶
-
Description: The
"segment-per-unit"option requests a hexagon-based SGE in the 10x genomics format. It requests the execution of RulesdgeAR_segment, alongside the prerequisite rules:sdgeAR_reformat,sdge2sdgeAR,dge2sdge,align,smatch,sbcd2chip, andfastq2sbcd. -
Rule Graph: